-
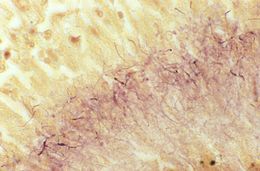
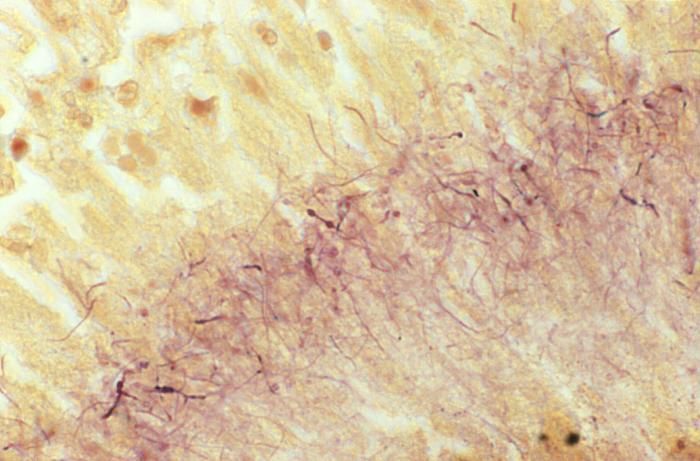
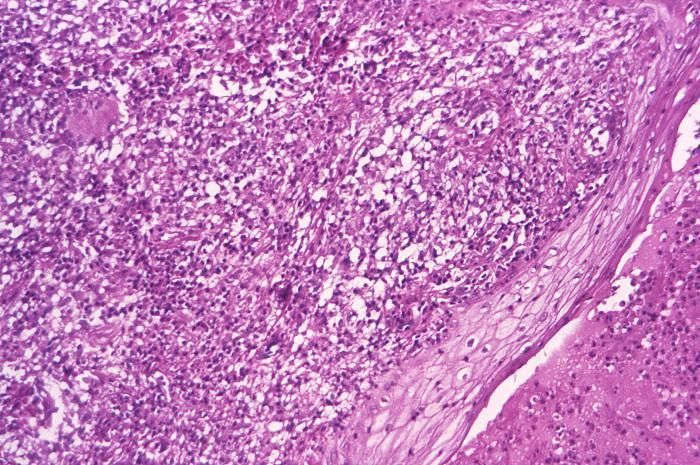
This Brown & Brenn-stained tissue specimen revealed the presence of Gram-positive Actinomadura madurae bacterial organisms.Created: 1974
-
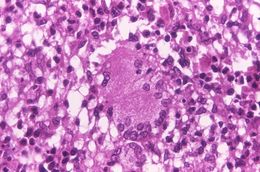
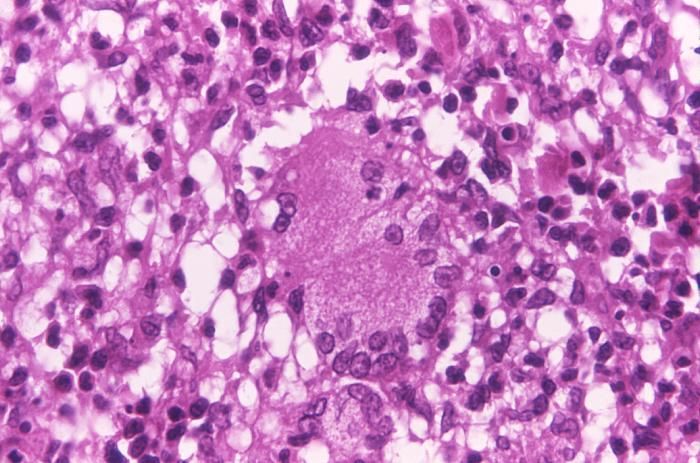
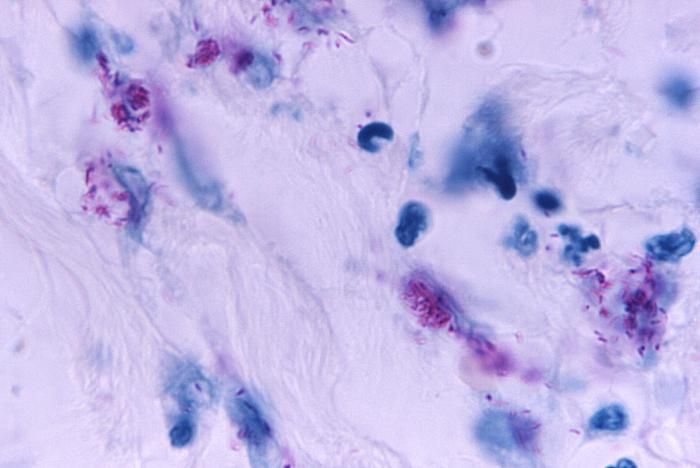
This light photomicrograph revealed some of the histopathologic cytoarchitectural characteristics seen in a mycobacterial skin infection.Created: 1972
-
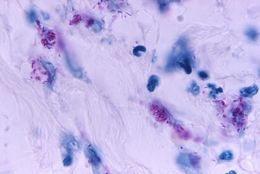
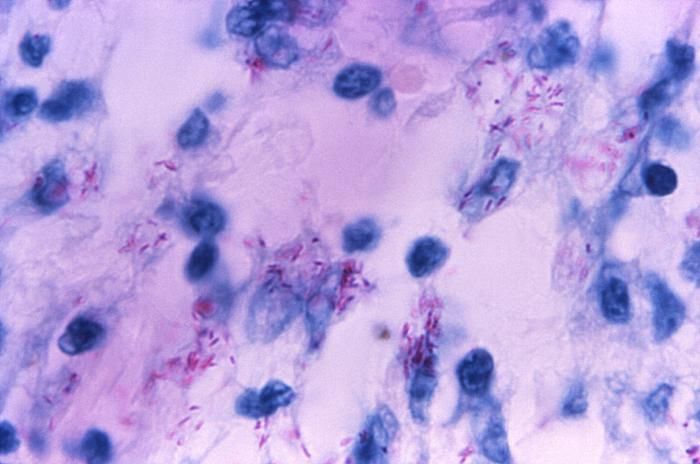
This light photomicrograph revealed some of the histopathologic cytoarchitectural characteristics seen in a mycobacterial skin infection.Created: 1972
-
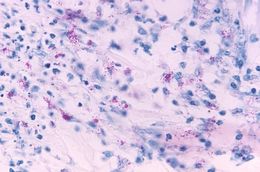
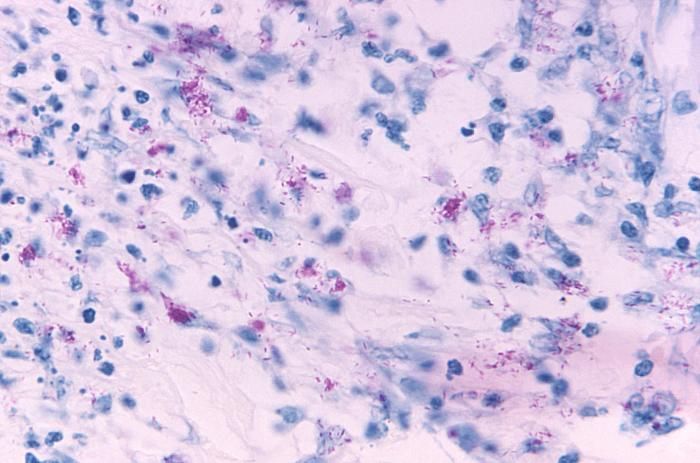
This light photomicrograph revealed some of the histopathologic cytoarchitectural characteristics seen in a mycobacterial skin infection.Created: 1972
-
This light photomicrograph revealed some of the histopathologic cytoarchitectural characteristics seen in a mycobacterial skin infection.Created: 1972
-
This light photomicrograph revealed some of the histopathologic cytoarchitectural characteristics seen in a mycobacterial skin infection.Created: 1972
-
This 1971 photomicrograph of a sputum smear, which had been stained using the Morse method of fluorescent acid-fast staining technique, revealed the presence of two Mycobacterium tuberculosis, rod-shaped bacteria, otherwise known as bacilli.Created: 1971
-
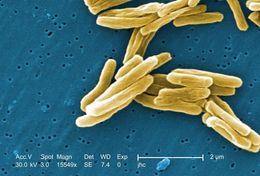
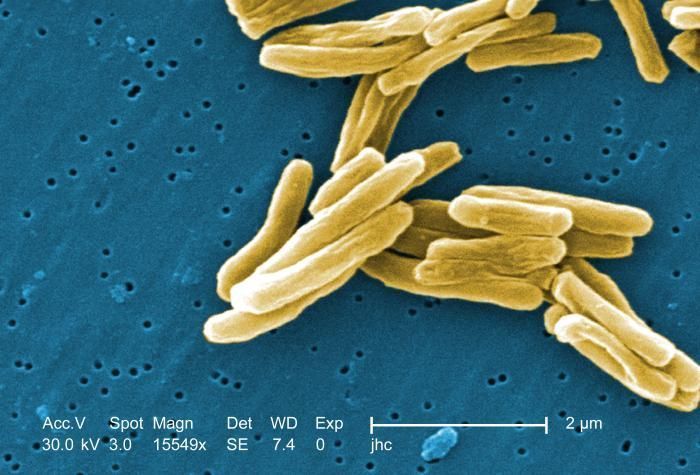
Under a high magnification of 15549x, this colorized scanning electron micrograph (SEM) depicted some of the ultrastructural details seen in the cell wall configuration of a number of Gram-positive Mycobacterium tuberculosis bacteria. As an obligate aerobic organism M. tuberculosis can only survive in an environment containing oxygen. This bacterium ranges in length between 2 - 4 microns, and a width between 0.2 - 0.5 microns. See PHIL 8438 for a black and white version of this image.Created: 2006
-
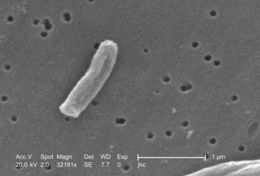
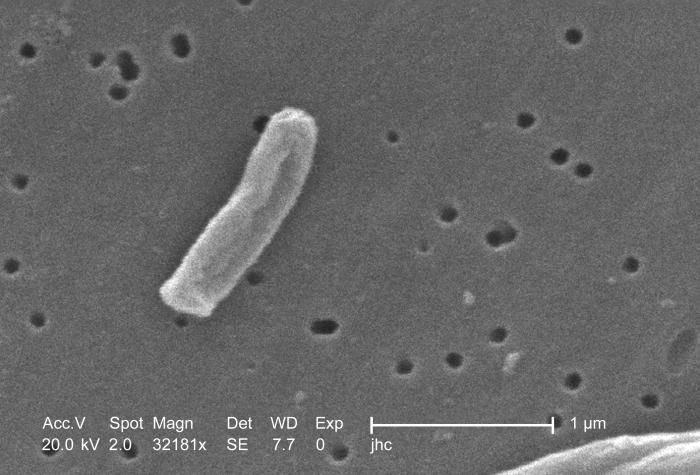
Under a very high magnification of 32181x, this scanning electron micrograph (SEM) depicted some of the ultrastructural details seen in the cell wall configuration of a Gram-positive Mycobacterium tuberculosis bacterium. As an obligate aerobic organism M. tuberculosis can only survive in an environment containing oxygen. This bacterium ranges in length between 2 - 4µm, and a width between 0.2 - 0.5µm.Created: 2006
-
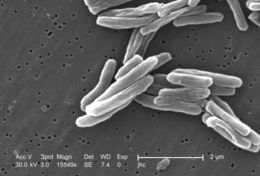
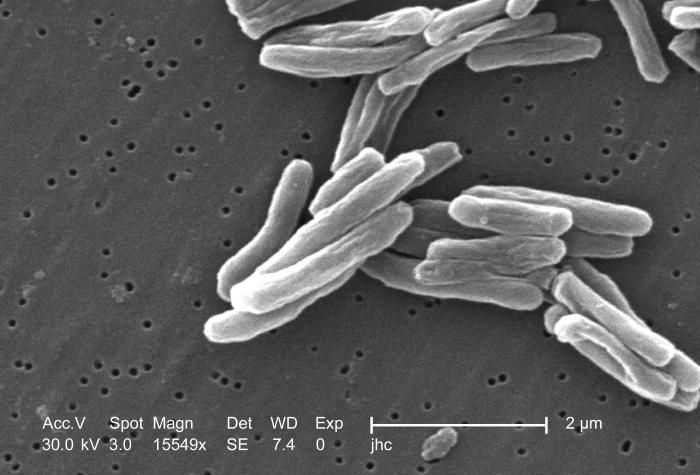
Under a high magnification of 15549x, this scanning electron micrograph (SEM) depicted some of the ultrastructural details seen in the cell wall configuration of a number of Gram-positive Mycobacterium tuberculosis bacteria. As an obligate aerobic organism M. tuberculosis can only survive in an environment containing oxygen. This bacterium ranges in length between 2 - 4 microns, and a width between 0.2 - 0.5 microns. See PHIL 9997 for a colorized version of this image.Created: 2006
-
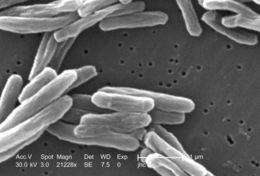
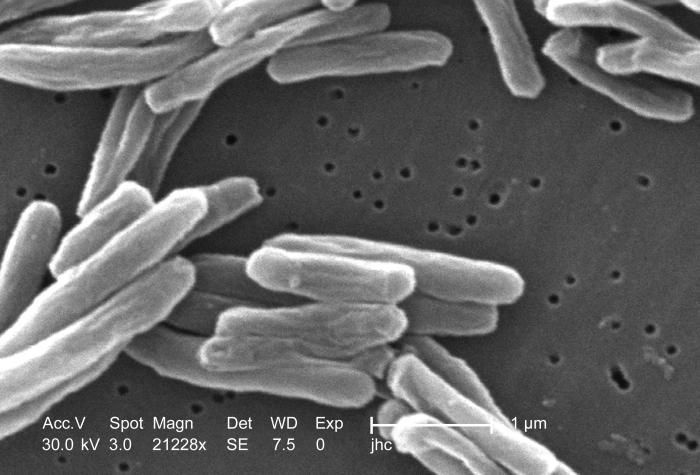
Under a high magnification of 21228x, this scanning electron micrograph (SEM) depicted some of the ultrastructural details seen in the cell wall configuration of a number of Gram-positive Mycobacterium tuberculosis bacteria. As an obligate aerobic organism M. tuberculosis can only survive in an environment containing oxygen. This bacterium ranges in length between 2 - 4µm, and a width between 0.2 - 0.5µm.Created: 2006
-
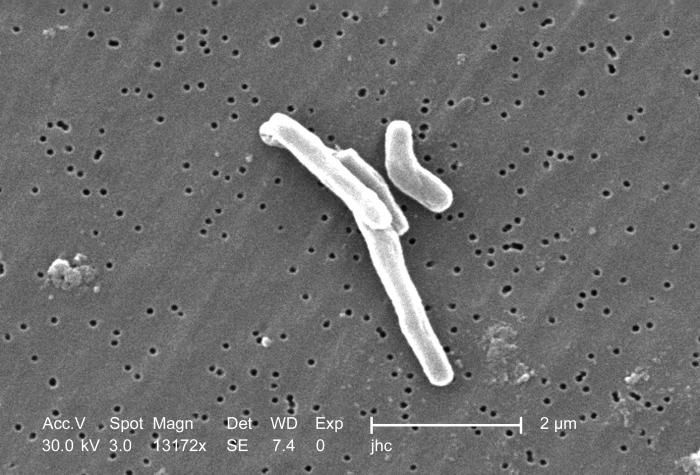
At a magnification of 13172x, this scanning electron micrograph (SEM) depicted a number of Gram-positive Mycobacterium tuberculosis bacteria. As an obligate aerobic organism M. tuberculosis can only survive in an environment containing oxygen. This bacterium ranges in length between 2 - 4µm, and a width between 0.2 - 0.5µm.Created: 2006
-
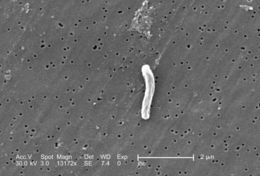
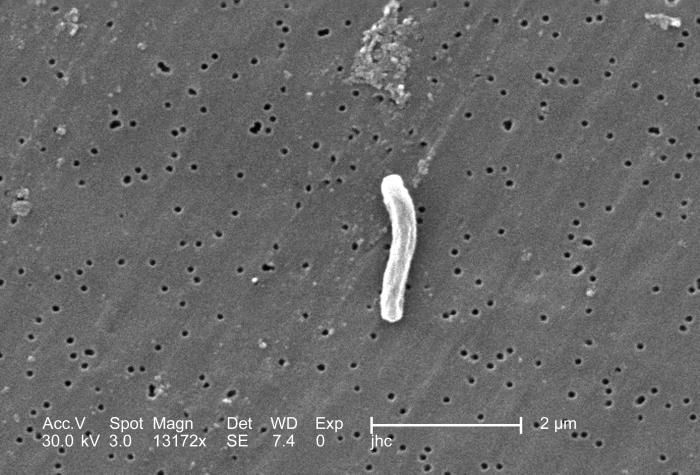
At a magnification of 13172x, this scanning electron micrograph (SEM) depicted a single Gram-positive Mycobacterium tuberculosis bacterium. As an obligate aerobic organism M. tuberculosis can only survive in an environment containing oxygen. This bacterium ranges in length between 2 - 4µm, and a width between 0.2 - 0.5µm.Created: 2006
-
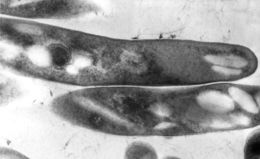
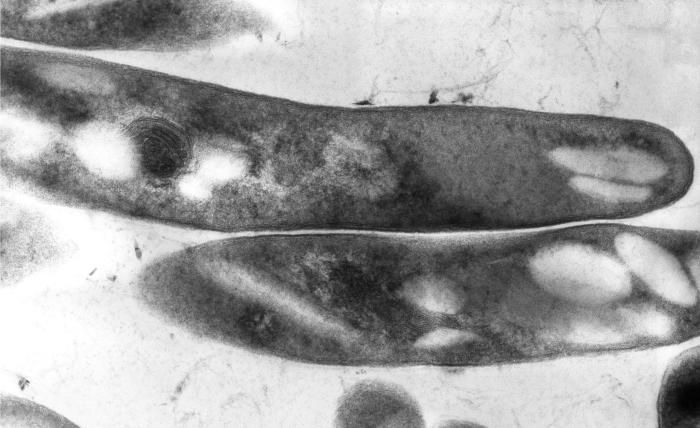
This thin section transmission electron micrograph (TEM) depicted the ultrastructural details displayed by a number of Gram-positive Mycobacterium tuberculosis bacilli, the causative agent for tuberculosis.Created:
-
-
-
-
-
-
-
-
-
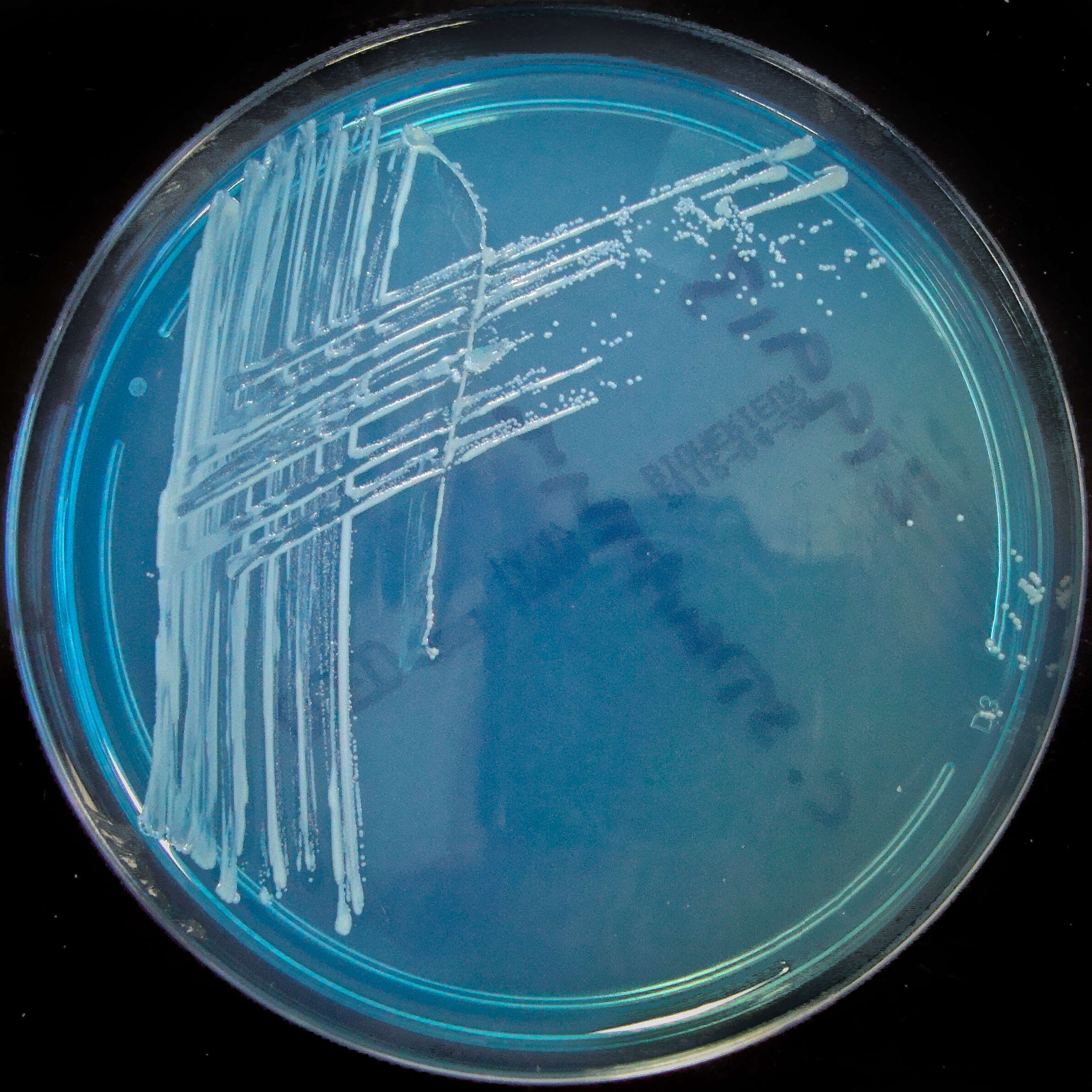
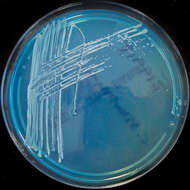
Summary.mw-parser-output table.commons-file-information-table,.mw-parser-output.fileinfotpl-type-information{border:1px solid #a2a9b1;background-color:#f8f9fa;padding:5px;font-size:95%;border-spacing:2px;box-sizing:border-box;margin:0;width:100%}.mw-parser-output table.commons-file-information-table>tbody>tr,.mw-parser-output.fileinfotpl-type-information>tbody>tr{vertical-align:top}.mw-parser-output table.commons-file-information-table>tbody>tr>td,.mw-parser-output table.commons-file-information-table>tbody>tr>th,.mw-parser-output.fileinfotpl-type-information>tbody>tr>td,.mw-parser-output.fileinfotpl-type-information>tbody>tr>th{padding:4px}.mw-parser-output.fileinfo-paramfield{background:#ccf;text-align:right;padding-right:0.4em;width:15%;font-weight:bold}.mw-parser-output.commons-file-information-table+table.commons-file-information-table,.mw-parser-output.commons-file-information-table+div.commons-file-information-table>table{border-top:0;padding-top:0;margin-top:-8px}@media only screen and (max-width:719px){.mw-parser-output table.commons-file-information-table,.mw-parser-output.commons-file-information-table.fileinfotpl-type-information{border-spacing:0;padding:0;word-break:break-word;width:100%!important}.mw-parser-output.commons-file-information-table>tbody,.mw-parser-output.fileinfotpl-type-information>tbody{display:block}.mw-parser-output.commons-file-information-table>tbody>tr>td,.mw-parser-output.commons-file-information-table>tbody>tr>th,.mw-parser-output.fileinfotpl-type-information>tbody>tr>td,.mw-parser-output.fileinfotpl-type-information>tbody>tr>th{padding:0.2em 0.4em;text-align:left;text-align:start}.mw-parser-output.commons-file-information-table>tbody>tr,.mw-parser-output.fileinfotpl-type-information>tbody>tr{display:flex;flex-direction:column}.mw-parser-output.commons-file-information-table+table.commons-file-information-table,.mw-parser-output.commons-file-information-table+div.commons-file-information-table>table{margin-top:-1px}.mw-parser-output.fileinfo-paramfield{box-sizing:border-box;flex:1 0 100%;width:100%}} Description: Isolated from a blood culture taken via a central line. Potential line coloniser. Date: 28 February 2012, 19:54. Source:
Corynebacterium striatum on C.L.E.D. agar. Author:
Nathan Reading from Halesowen, UK. Other versions:
Detail.
-
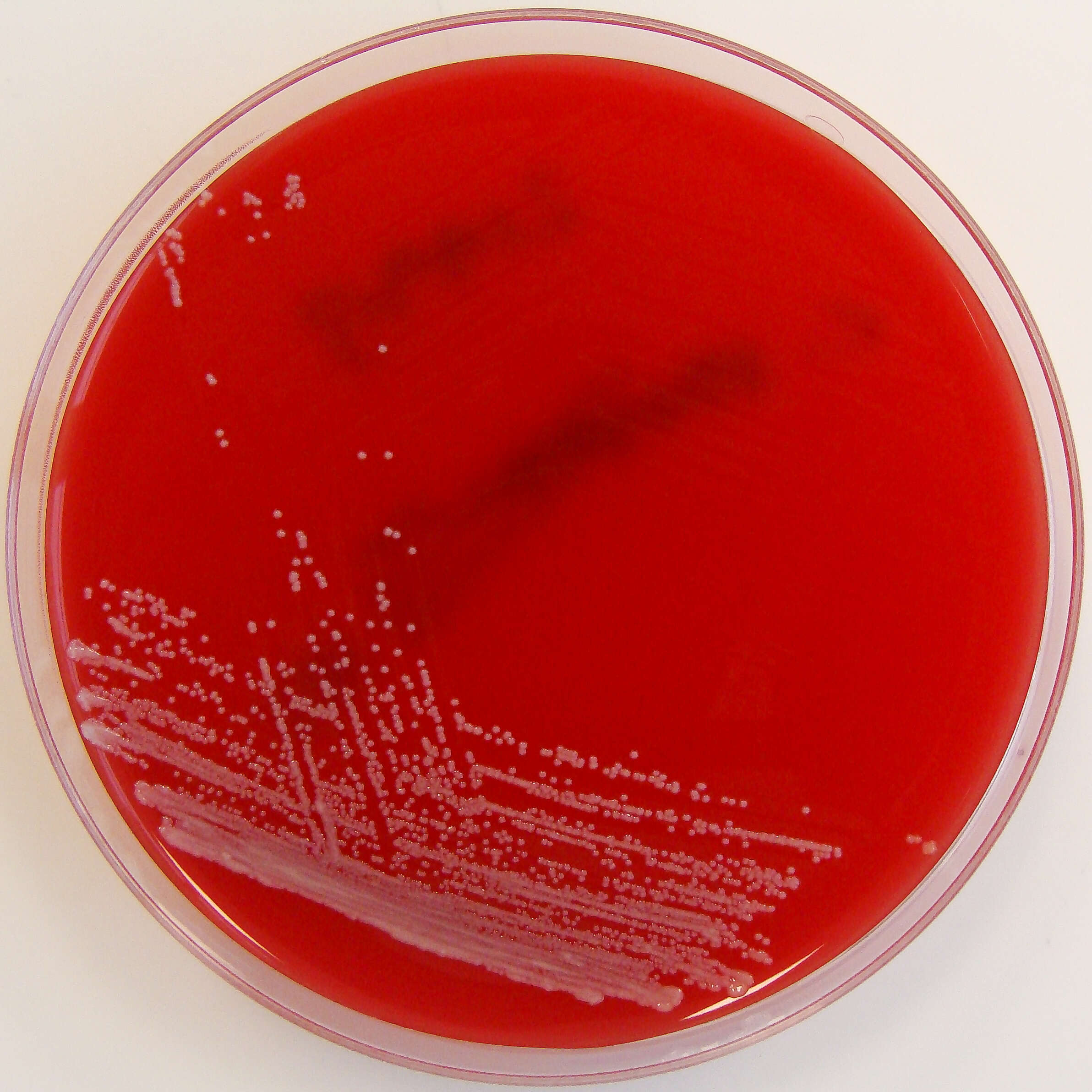
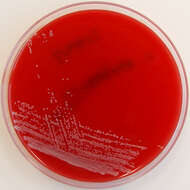
Summary.mw-parser-output table.commons-file-information-table,.mw-parser-output.fileinfotpl-type-information{border:1px solid #a2a9b1;background-color:#f8f9fa;padding:5px;font-size:95%;border-spacing:2px;box-sizing:border-box;margin:0;width:100%}.mw-parser-output table.commons-file-information-table>tbody>tr,.mw-parser-output.fileinfotpl-type-information>tbody>tr{vertical-align:top}.mw-parser-output table.commons-file-information-table>tbody>tr>td,.mw-parser-output table.commons-file-information-table>tbody>tr>th,.mw-parser-output.fileinfotpl-type-information>tbody>tr>td,.mw-parser-output.fileinfotpl-type-information>tbody>tr>th{padding:4px}.mw-parser-output.fileinfo-paramfield{background:#ccf;text-align:right;padding-right:0.4em;width:15%;font-weight:bold}.mw-parser-output.commons-file-information-table+table.commons-file-information-table,.mw-parser-output.commons-file-information-table+div.commons-file-information-table>table{border-top:0;padding-top:0;margin-top:-8px}@media only screen and (max-width:719px){.mw-parser-output table.commons-file-information-table,.mw-parser-output.commons-file-information-table.fileinfotpl-type-information{border-spacing:0;padding:0;word-break:break-word;width:100%!important}.mw-parser-output.commons-file-information-table>tbody,.mw-parser-output.fileinfotpl-type-information>tbody{display:block}.mw-parser-output.commons-file-information-table>tbody>tr>td,.mw-parser-output.commons-file-information-table>tbody>tr>th,.mw-parser-output.fileinfotpl-type-information>tbody>tr>td,.mw-parser-output.fileinfotpl-type-information>tbody>tr>th{padding:0.2em 0.4em;text-align:left;text-align:start}.mw-parser-output.commons-file-information-table>tbody>tr,.mw-parser-output.fileinfotpl-type-information>tbody>tr{display:flex;flex-direction:column}.mw-parser-output.commons-file-information-table+table.commons-file-information-table,.mw-parser-output.commons-file-information-table+div.commons-file-information-table>table{margin-top:-1px}.mw-parser-output.fileinfo-paramfield{box-sizing:border-box;flex:1 0 100%;width:100%}} Description: Isolated from a blood culture taken via a central line. Potential line coloniser. Date: 28 February 2012, 19:51. Source:
Corynebacterium striatum on Columbia Horse Blood Agar. Author:
Nathan Reading from Halesowen, UK. Other versions:
Detail.